在处理图形数据时,我遇到了以下子任务:
我需要以 COO 格式构建图连接矩阵 对于具有来自“边界”索引数组的几个完全连接组件的图形。
例如,给定数组
borders = [0, 2, 5]
coo_matrix = [[0, 0, 1, 1, 2, 2, 2, 3, 3, 3, 4, 4, 4],
[0, 1, 0, 1, 2, 3, 4, 2, 3, 4, 2, 3, 4]].
我想出了以下算法,但我怀疑性能可以提高:
import numpy as np
def get_coo(borders):
edge_list = []
for s, e in zip(borders, borders[1:]):
# create fully-connected subgraph
arr = np.arange(s, e)
t = np.array(np.meshgrid(arr, arr)).T.reshape(-1, 2)
t = t.T
edge_list.append(t)
edge_list = np.concatenate(edge_list, axis=1)
return edge_list
有没有人有任何想法?
最佳答案
由于您的目标是比现有解决方案更快的解决方案,您可以探索 itertools为了有效地解决这个问题。这种方法基准测试 约 25 次 比在更大的 border 上测试的当前方法更快列表。
import numpy as np
from itertools import product, chain
def get_coo(borders):
edges = chain(*[product(range(*i),repeat=2) for i in zip(borders, borders[1:])])
return list(edges)
output = get_coo(borders)
## NOTE: Remember to can convert to array and Transpose for all approaches below to get Coo format.
np.array(output).T
array([[0, 0, 1, 1, 2, 2, 2, 3, 3, 3, 4, 4, 4],
[0, 1, 0, 1, 2, 3, 4, 2, 3, 4, 2, 3, 4]])
替代方法和基准:
Note: These have been benchmarked on your current small list as well as on a larger border list as generated by
borders = np.arange(300)[np.random.randint(0,2,(300,),dtype=bool)]
完全图的不相交并集
您要做的基本上是组合不相交的完整图。此类图的邻接矩阵沿其对角线具有选择项的完整连接。您可以使用
networkx来解决这些。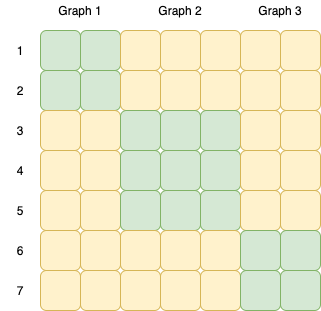
虽然比您当前的解决方案慢,但您会发现处理这些图形对象比使用 NumPy 来表示图形要容易得多,而且返回也大得多。
方法一:
i i*i nx.disjoint_union_all 组合图形import numpy as np
import networkx as nx
def get_coo(borders):
graphs = [nx.from_numpy_matrix(np.ones((i,i))).to_directed() for i in np.diff(borders)]
edges = nx.disjoint_union_all(graphs).edges()
return edges
%timeit get_coo(borders)
#Small- 277 µs ± 33.5 µs per loop (mean ± std. dev. of 7 runs, 1000 loops each)
#Large- 300 ms ± 36.1 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
borders 的滚动 2 克元组使用 zip nx.complete_graph使用 ranges(start, end)来自这些元组 nx.disjoint_union_all 组合图形import numpy as np
import networkx as nx
def get_coo(borders):
graphs = [nx.complete_graph(range(*i),nx.DiGraph()) for i in zip(borders, borders[1:])]
edges = nx.disjoint_union_all(graphs).edges()
return edges
%timeit get_coo(borders)
#Small- 116 µs ± 13.4 µs per loop (mean ± std. dev. of 7 runs, 10000 loops each)
#Large- 207 ms ± 35.5 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
使用
itertools.product 方法三:borders 的滚动 2 克元组使用 zip itertools.product每个子图的完全连接边列表 itertools.chain “附加”两个迭代器 import numpy as np
from itertools import product, chain
def get_coo(borders):
edges = chain(*[product(range(*i),repeat=2) for i in zip(borders, borders[1:])])
return list(edges)
%timeit get_coo(borders)
#Small- 3.91 µs ± 787 ns per loop (mean ± std. dev. of 7 runs, 100000 loops each)
#Large- 183 µs ± 21.7 µs per loop (mean ± std. dev. of 7 runs, 1000 loops each)
您当前的方法 - 基准
def get_coo(borders):
edge_list = []
for s, e in zip(borders, borders[1:]):
# create fully-connected subgraph
arr = np.arange(s, e)
t = np.array(np.meshgrid(arr, arr)).T.reshape(-1, 2)
t = t.T
edge_list.append(t)
edge_list = np.concatenate(edge_list, axis=1)
return edge_list
%timeit get_coo(borders)
#Small- 95.1 µs ± 10.8 µs per loop (mean ± std. dev. of 7 runs, 10000 loops each)
#Large- 3.91 ms ± 962 µs per loop (mean ± std. dev. of 7 runs, 100 loops each)
关于python - 以 COO 格式构建图连接矩阵,我们在Stack Overflow上找到一个类似的问题: https://stackoverflow.com/questions/69260592/